 
Their results showed that the SARS-CoV-2 spike protein activated the natural immune response in heart muscle cells and damaged the heart, but the NL63 spike protein did not. They studied the potential ability to cause heart disease of both SARS-CoV-2 spike protein and the NL63 spike protein. HCoV-NL63 is a coronavirus that infects the respiratory system without causing cardiac injury, although its spike protein also uses ACE2 to mediate virus entry. The researchers studied whether the SARS-CoV-2 spike protein activates the natural immune response in heart muscle cells. Activation of the body's immune response is essential for fighting against virus infection however, this may also impair heart muscle cell function and even lead to cell death and heart failure," Lin said. "Host natural immunity is the first line of defense against pathogen invasion, and heart muscle cells have their own natural immune machinery. That's why it's important to get vaccinated and prevent this disease." "Our data show that the spike protein from SARS-CoV-2 causes heart muscle damage. What we suspect is that the spike protein has unknown pathological roles," said Zhiqiang Lin, Ph.D., lead author of the study and an assistant professor at the Masonic Medical Research Institute in Utica, New York. "It's already known from the clinical side that COVID-19 infection can induce heart injury, however, what we don't know is the mechanistic details of how this occurs. In addition to infecting the lungs, the virus can also spread to other organs leading to more damage to the body, severe infection and, among some people, death. The spike protein facilitates virus entry into healthy cells, which is the first step in infection. Spike proteins latch onto receptors known as angiotensin-converting enzyme 2 (ACE2) on target cells. These studies will allow us to determine common principals that can be applied to all CoVs, moving the field forward with these innovative studies will provide critical knowledge about CoV entry and tropism needed to safeguard human health from an emerging pathogen likely to cause severe outbreaks, and for which few or no medical countermeasures exist.The spike protein is found on the surface of SARS-CoV-2, the virus that causes COVID-19. 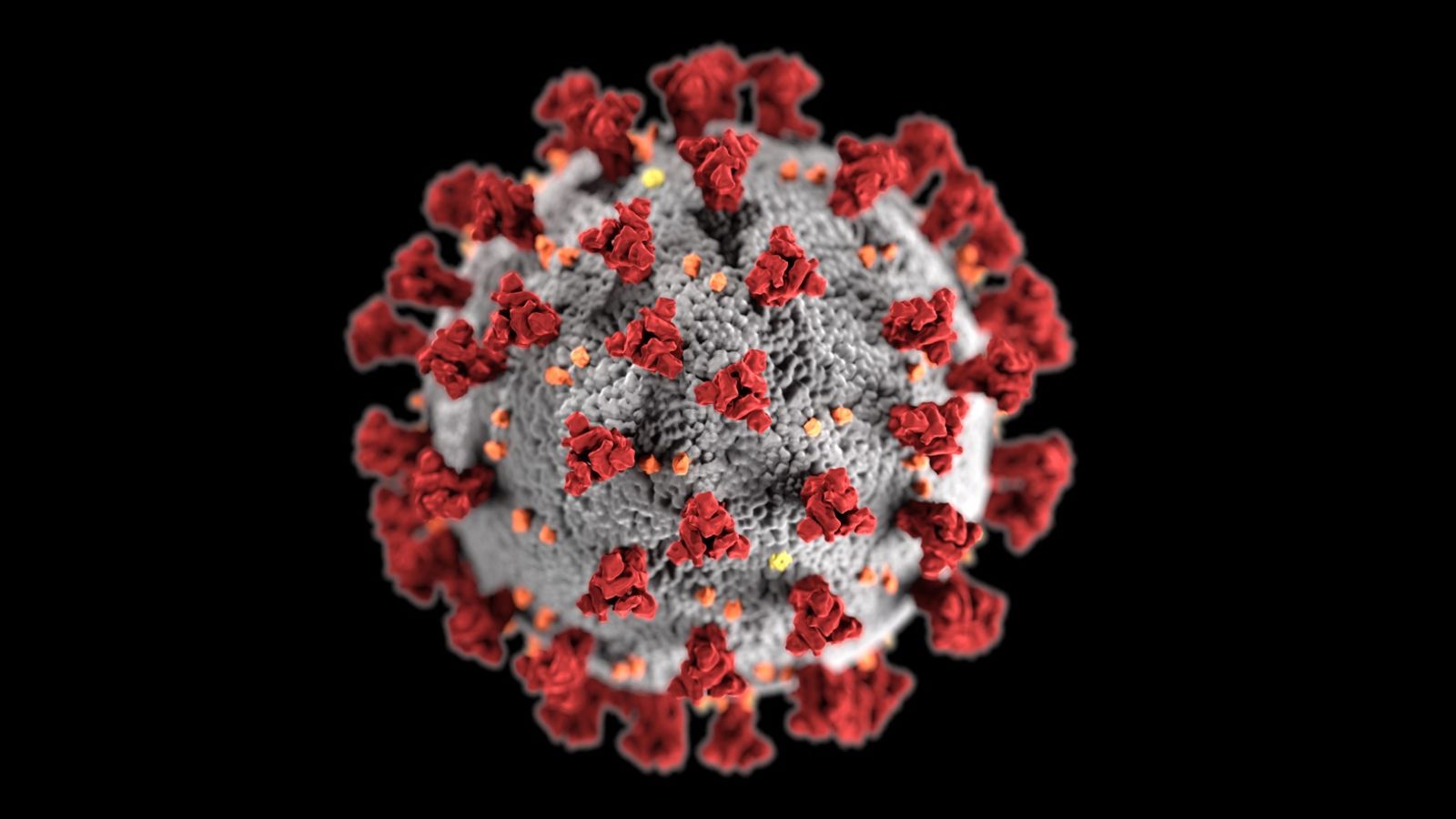
receptor binding, protease availability and the local ionic environment. These approaches will reveal how structure and function vary depending on the key activators of S i.e. We will use state-of-the-art spectroscopy and an innovative single particle tracking technique to study S protein fusion peptide function, and combine these with in vivo infectivity studies, including at BS元, will allow a complete picture of CoV fusion activation. We propose an integrated biophysical, biochemical, and in vivo approach to study the unique cleavage-activated regulation of CoV S protein, using Middle East respiratory syndrome coronavirus (MERS-CoV) and severe acute respiratory syndrome coronavirus (SARS-CoV) as primary models. This information is critical to understanding viral pathogenesis and CoV emergence into the human population. Despite the recent availability of S structures in their prefusion state, there remains a very limited mechanistic understanding of membrane fusion for the CoV family, or any structural information to correlate structural biology aspects of S to its function in membrane fusion. Thus fusion peptide activity is likely controlled by reorganization of the fusion platform, based on both hydrophobic (i.e. Thus, there are notable differences between CoV S and most other class I fusion proteins including: 1) that the proteolytic events liberating the fusion peptide are diverse, and 2) that the fusion peptide itself is atypical in sequence compared to other fusion peptides, containing a mixture of important hydrophobic and negatively charged residues, and may represent a larger than normal fusion “platform” instead of a defined “peptide”. 
We have pioneered the concept that that the proteolytic cleavage events in S that lead to membrane fusion occur both at the interface of the receptor binding (S1) and fusion (S2) domains (called S1/S2), as well as adjacent to a structurally and functionally novel fusion peptide within S2 (called S2’). This is exploited by these viruses in their diverse entry pathways and is a primary determinant of viral tropism. 
Coronavirus fusion spike protein (CoV S) is a complex biomolecular machine that has a novel fusion peptide with has a great deal of inherent flexibility in its fusion reaction. Both binding and fusion are mediated by a specific viral “spike” protein that is typically primed for fusion activation by proteolytic cleavage to expose the fusion peptide. Enveloped viruses access their host cells by binding to receptors on the plasma membrane and then undergoing fusion with the host membrane. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
